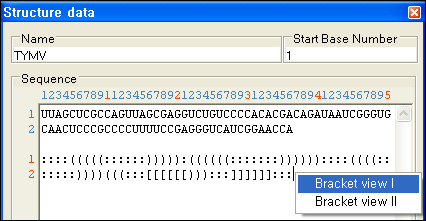
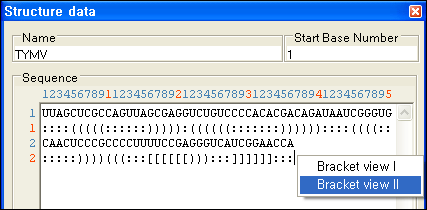
Paired Positions
If a user knows the stem positions only and does not know how to
represent the structure in bracket view, he can simply provide the stem
positions in the following format, which is also used in PseudoBase. Given stem
positions, PseudoViewer3 automatically generates the structure data in bracket
view and visualize the structure.
|
# <RNA name>
// optional line
<starting base number> //optional;
if the starting base number is not specified, it is 1 by default.
<base sequence >
Positions paired.
<Starting position of the stem opening part>-<ending
position of the stem opening part>;
<starting position of the stem-closing part>-<ending
position of the stem closing part>
|
# Hepatitis delta virus
(pseudoknot only)
688
GGCCGGCAUGGUCCCAGCCUCCUCGCUGGCGCCGGCUGGGCAA
CAUUCCGAGGGGACCGUCCCCUCGGUAAUGGCGAAUGGGACC
Positions paired.
688-694; 718-724
697-703; 766-772
704-706; 715-717
708-709; 725-726
731-734; 757-760
735-743; 747-755
|
Multiple sequences
If some part of sequence is omitted, user can use the input format follows.
|
# <RNA name >
// optional line
<starting base number> //
optional; if the starting base number is not specified, it is 1 by default. <base
sequence> //alternates with
<matching parentheses and brackets>
<starting base number> //
optional; if this is omitted, the starting number is 1 by default.
/<base
sequence> //alternates with
/<matching parentheses and brackets>
|
# PKB128 (Berne virus)
1045
UUUAAACUGUUGAGAGGUGCCUGGAGCGCCUGCAGGCAUCUCUGUU
............(((((((((((....[[[[[)))))))))))...
1150
/UAUAAUGCAGGCA
/.......]]]]].
|
bpseq format
|
ct format
|
Tertiary interaction data |
| Tertiary interactions other than pseudoknots are represented in a pair of
matching braces (ˇ®{}ˇŻ) in the structure data, and are displayed in dotted lines
in the final structure drawing. It helps represent other tertiary interactions
than pseudoknots. |